The hypothalamus was dissected from 3 chicken embryos on embryonic day 19.
Lab protocols from 10x Genomics were used to process the brain tissue and create gene expression and ATAC (assay of accessible chromatin) libraries, after which sequencing was performed using the Illumina NovaSeq 6000 system.
Cellranger ARC, a pipeline developed by 10x Genomics, was used to map all the reads to the chicken genome and transcriptome, and to generate basic summary metrics.
Quality control
Ambient RNA is one of the data quality problems that occur with single-cell data, because some of the cells lyse before they should. This causes RNA to just float around in the cell solution. In our case, we only wanted to collect data from the nuclei. Therefore, we can use statistics that give insights into the amount of data originating from cytoplasmic RNA to estimate the amount of ambient RNA.
Enrichment around the transcription start site
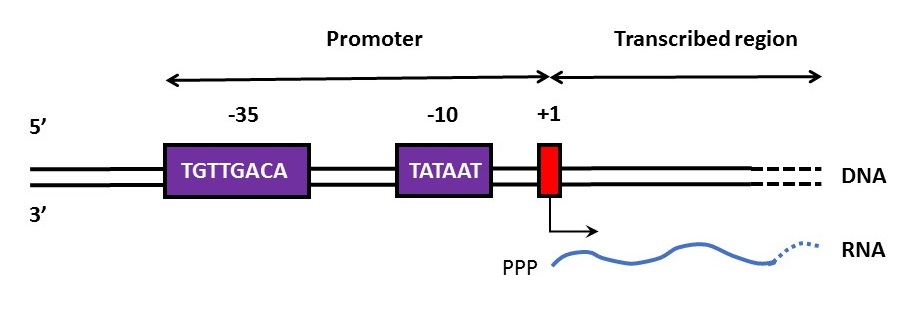
To study accessibility of chromatin, the open chromatin is cut by a transposase. When the region around the transcription start site is open, we expect a relatively high number of cuts in this region. Graphs were made to be able to see the distribution of relative number of cuts.
ATAC clusters
Clustering methods can divide the cells that we have into distinct groups. In this way we can try to match each cell with the right cell type, to then study each cell type population by itself. Clustering was done by making a 2-D-t-SNE projection of PCA components.